Visualization with inquanto-nglview
It is often useful to visualize certain molecular data. For this purpose, InQuanto uses an extension to interface with the NGLview nglview package. The inquanto-nglview extension is focused on providing some basic visualizing utilities to the user. The return types from the functions are widgets which can be viewed in a jupyter notebook. The functionality includes visualizing molecular
structures, fragmentation patterns defined in the Geometry object and molecular orbitals.
Visualizing Structures
VisualizerNGL is initialized by an InQuanto Geometry object specifying molecular, or unit cell, geometry. The molecule can then be visualized by calling the .visualize_molecule() in the last line of a notebook cell. See below for a short example.
[1]:
# pip install inquanto-nglview --index https://...
from inquanto.geometries import GeometryMolecular
from inquanto.extensions.nglview import VisualizerNGL
xyz = [
['C', [ 0.0000000, 1.4113170, 0.0000000]],
['C', [ 1.2222370, 0.7056590, 0.0000000]],
['C', [ 1.2222370, -0.7056590, 0.0000000]],
['C', [ 0.0000000, -1.4113170, 0.0000000]],
['C', [-1.2222370, -0.7056590, 0.0000000]],
['C', [-1.2222370, 0.7056590, 0.0000000]],
['H', [ 0.0000000, 2.5070120, 0.0000000]],
['H', [ 2.1711360, 1.2535060, 0.0000000]],
['H', [ 2.1711360, -1.2535060, 0.0000000]],
['H', [ 0.0000000, -2.5070120, 0.0000000]],
['H', [-2.1711360, -1.2535060, 0.0000000]],
['H', [-2.1711360, 1.2535060, 0.0000000]]
]
g = GeometryMolecular(xyz)
visualizer = VisualizerNGL(g)
[2]:
#visualizer.visualize_molecule(atom_labels="index").display(gui=True)
#markdown cell below is a presaved image
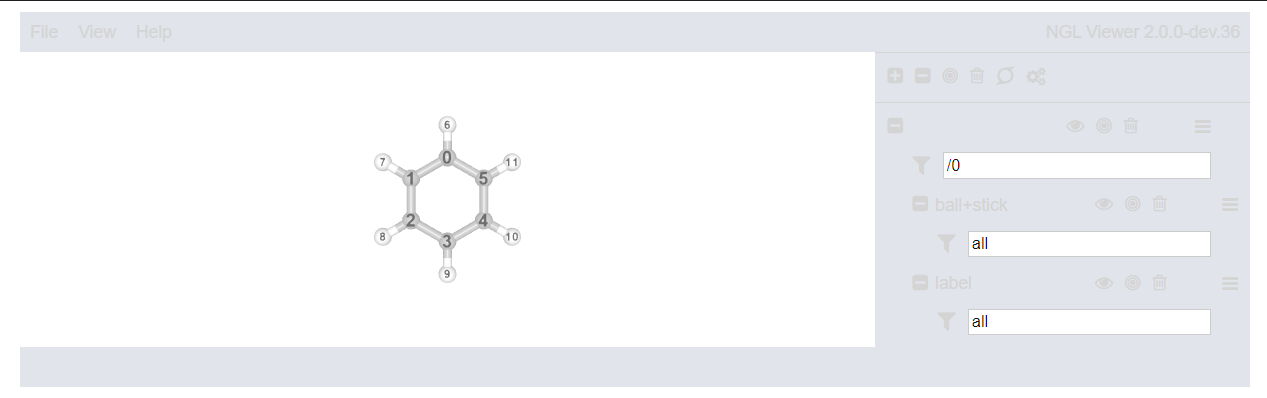
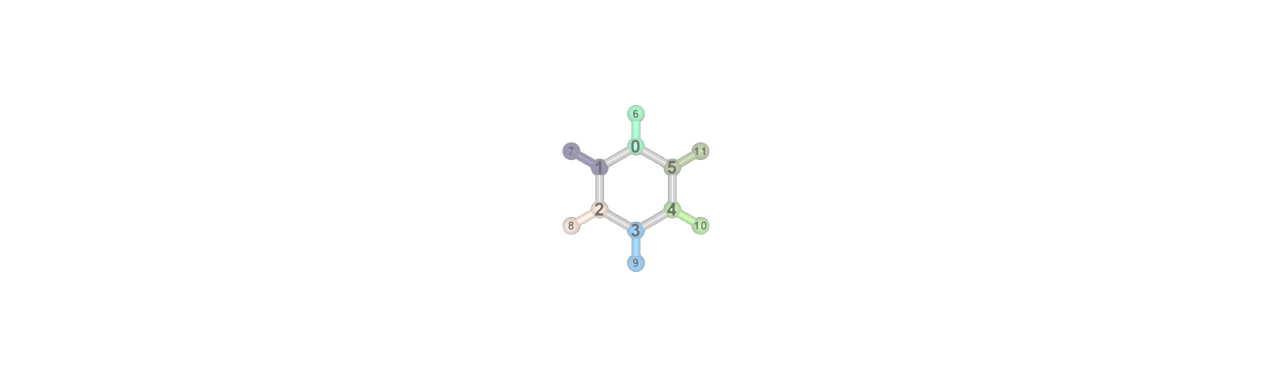
Visualizing Fragments
There are several fragmentation methods available in inquanto, to visualize a fragmentation defined in a Geometry object, call the visualize_fragmentation() method as illustrated in the code-snippet below.
[3]:
g.set_groups(
"fragments",
{
"ch1": [0, 6],
"ch2": [5, 11],
"ch3": [4, 10],
"ch4": [3, 9],
"ch5": [2, 8],
"ch6": [1, 7]
}
)
[4]:
#visualizer.visualize_fragmentation("fragments", atom_labels="index")
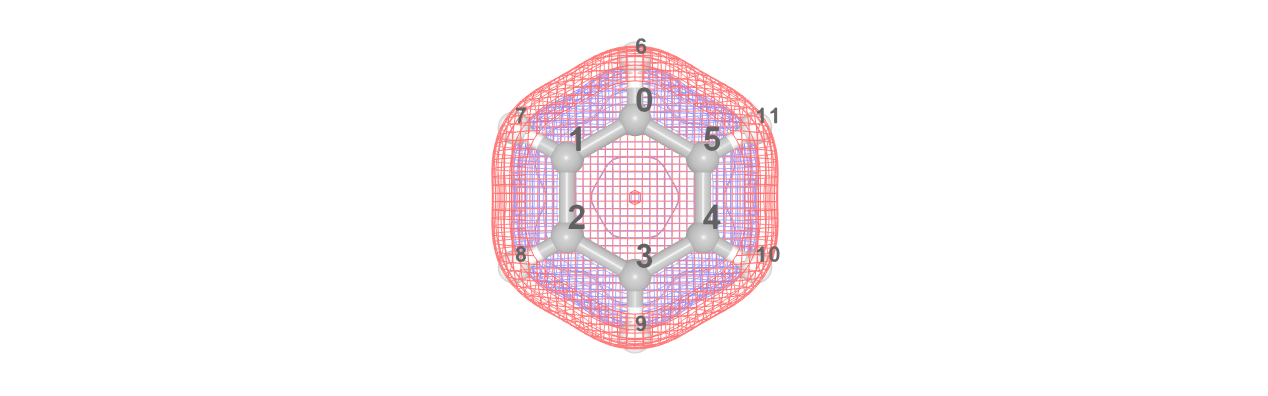
Visualizing Orbitals
To aid in active space selection, it is also possible to view the molecular orbitals of a system if the appropriate .cube strings are available.
[5]:
from inquanto.extensions.pyscf import ChemistryDriverPySCFMolecularRHF
driver = ChemistryDriverPySCFMolecularRHF(geometry=g.xyz, basis="sto3g")
# runs HF and returns orbitals
cube_orbitals=driver.get_cube_orbitals()
ngl_mos = [visualizer.visualize_orbitals(orb, atom_labels="index") for i, orb in enumerate(cube_orbitals)]
[6]:
#ngl_mos[16]